TeachOpenCADD¶
This website is free and open to all users and there is no login requirement!
Open source programming packages for cheminformatics and structural bioinformatics are powerful tools to build modular, reproducible, and reusable pipelines for computer-aided drug design (CADD). While documentation for such tools is available, only few freely accessible examples teach underlying concepts focused on CADD applications, addressing especially users new to the field.
TeachOpenCADD is a teaching platform developed by students for students, which provides teaching material for central CADD topics. Since we cover both the theoretical as well as practical aspect of these topics, the platform addresses students and researchers with a biological/chemical as well as a computational background.
For each topic, an interactive Jupyter Notebook is offered, using open source packages such as the Python packages rdkit, pypdb, biopandas, nglview, and mdanalysis. Topics are continuously expanded and open for contributions from the community. Beyond their teaching purpose, the TeachOpenCADD material can serve as starting point for users’ project-directed modifications and extensions.
New edition: we have extended the TeachOpenCADD platform with 6 notebooks introducing deep learning and its application to CADD related topics.
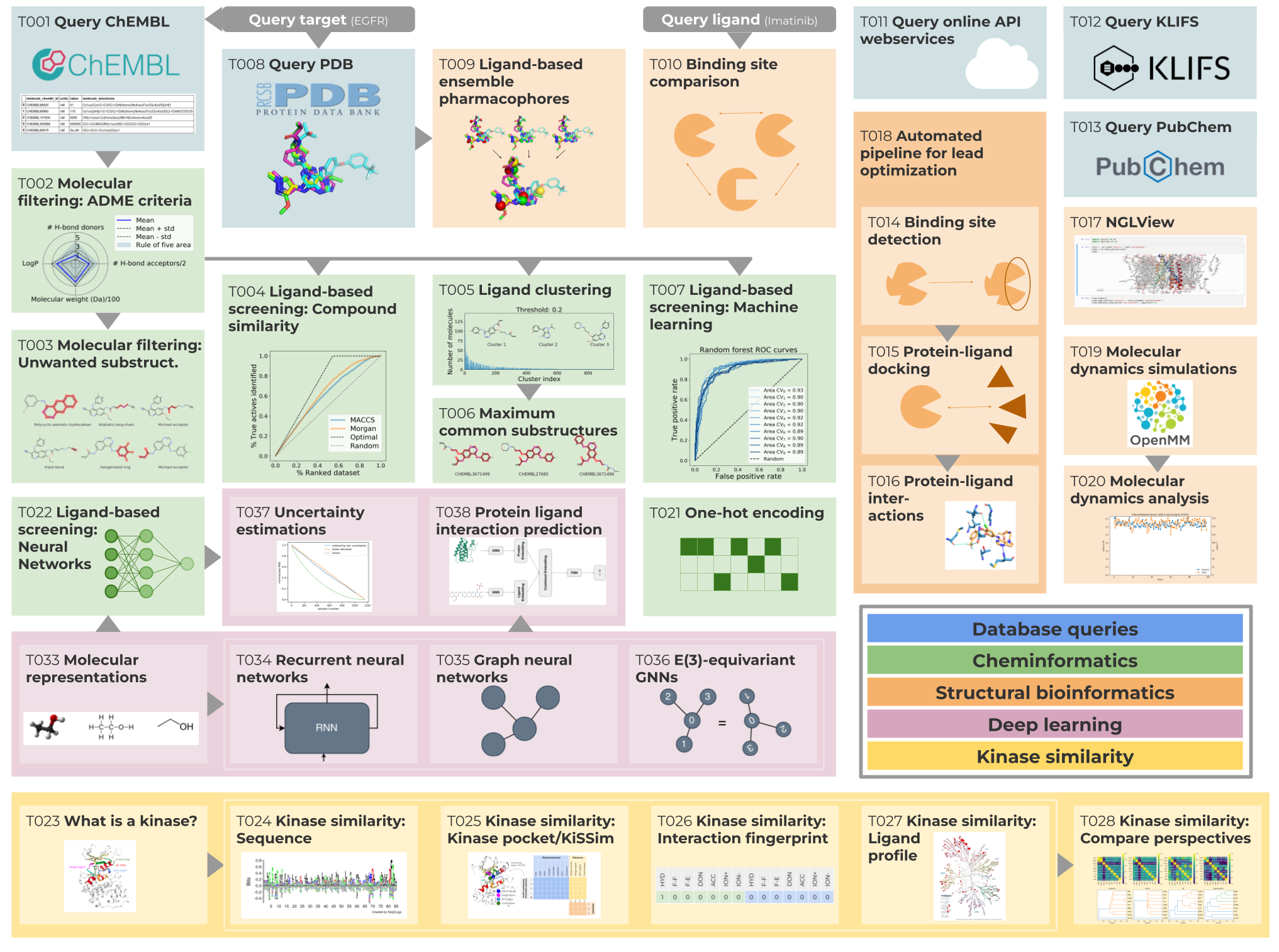
Figure adapted from Figure 1 in the TeachOpenCADD publication
(D. Sydow et al., J. Cheminformatics, 2019).
Table of contents¶
Our talktorials
User guide
Development
About TeachOpenCADD
External resources